RNA-Seq Quantification of Hepatic Drug Processing Genes in Germ-Free Mice | Drug Metabolism & Disposition
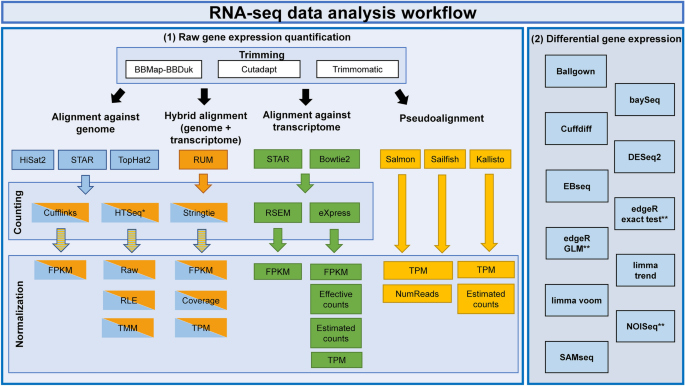
Systematic comparison and assessment of RNA-seq procedures for gene expression quantitative analysis | Scientific Reports
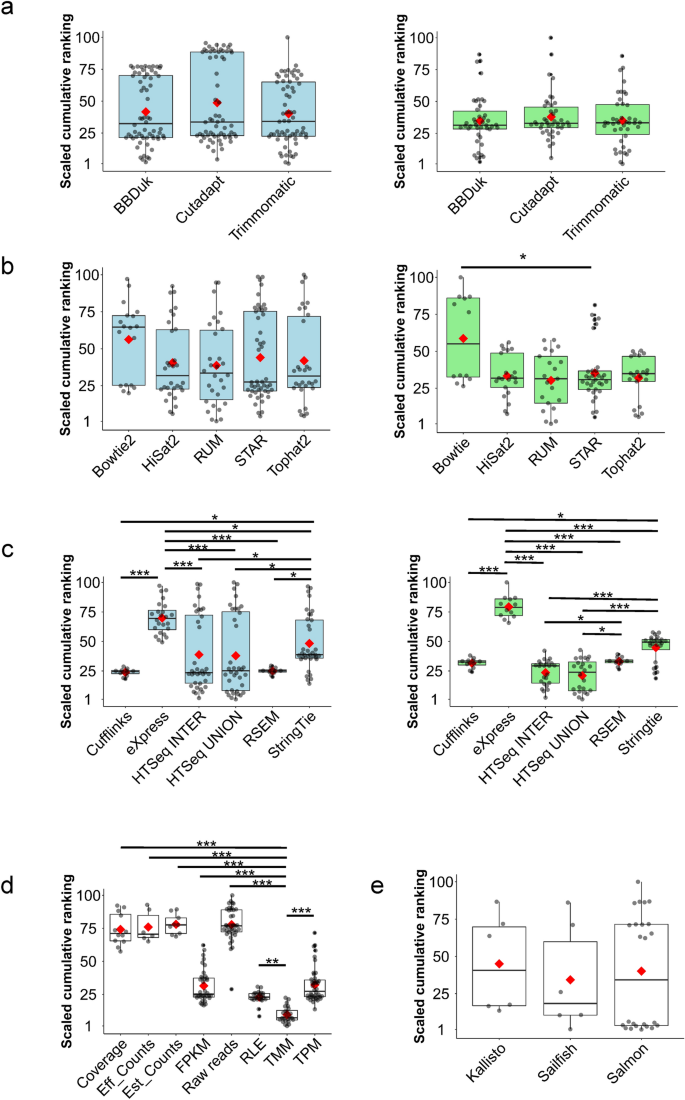
Systematic comparison and assessment of RNA-seq procedures for gene expression quantitative analysis | Scientific Reports
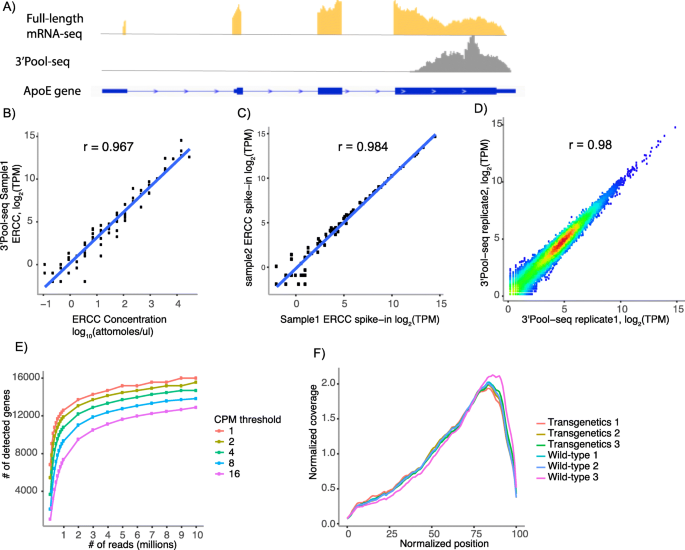
3'Pool-seq: an optimized cost-efficient and scalable method of whole-transcriptome gene expression profiling | BMC Genomics | Full Text

Quantification of transgene expression in GSH AAVS1 with a novel CRISPR/Cas9-based approach reveals high transcriptional variation: Molecular Therapy - Methods & Clinical Development

Detection and Quantification of GPCR mRNA: An Assessment and Implications of Data from High-Content Methods | ACS Omega

Quantification of differential gene expression by multiplexed targeted resequencing of cDNA | Nature Communications
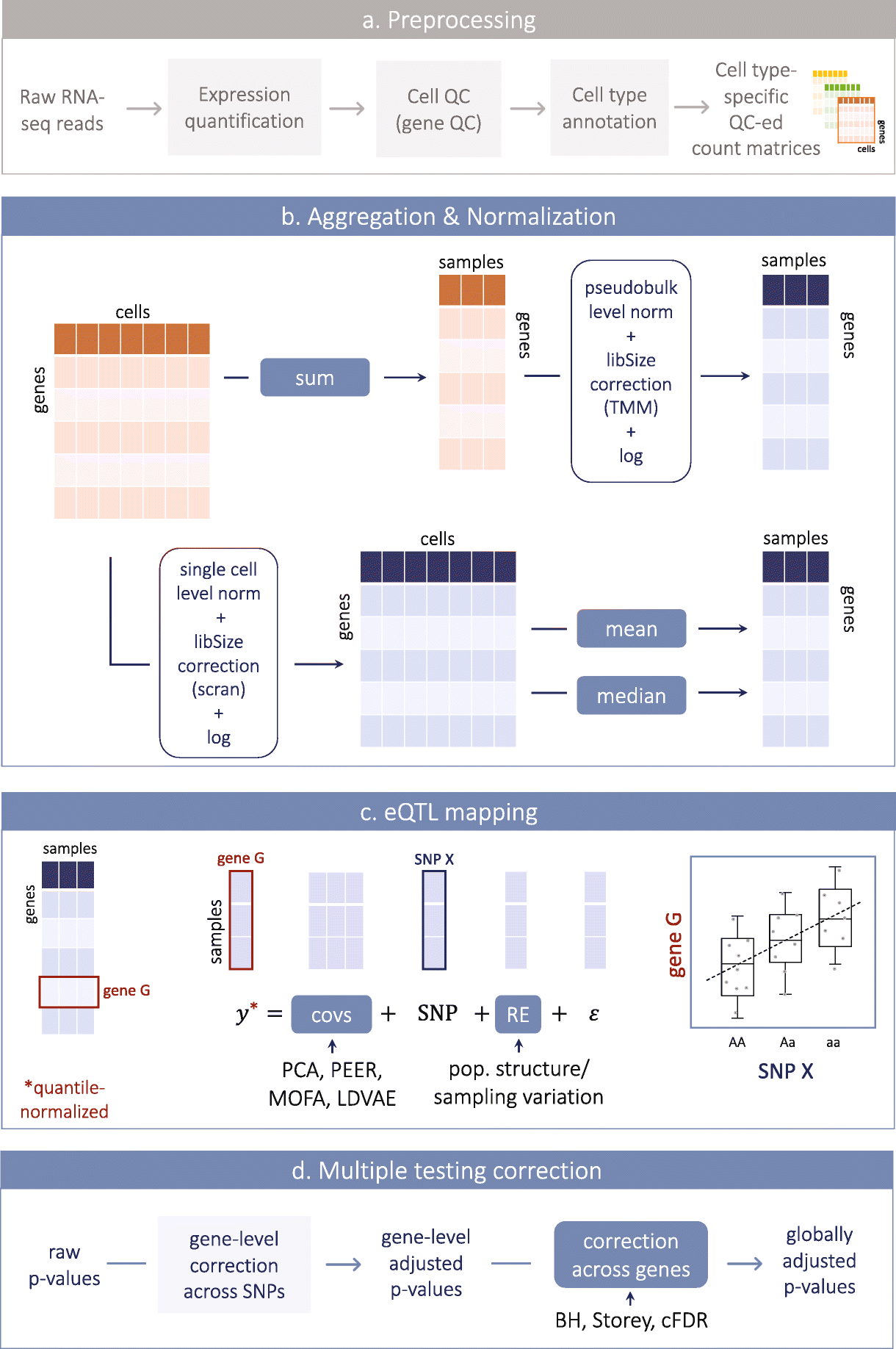
Optimizing expression quantitative trait locus mapping workflows for single-cell studies | Genome Biology | Full Text